Introduction to Lipidomics and Systems Biology
Lipids are fundamental biological molecules that participate in nearly every aspect of cellular organization, metabolism, signaling, and physiological regulation. Traditionally, lipids were viewed primarily as structural membrane components and energy storage molecules. However, advances in analytical chemistry, molecular biology, and computational biology have radically transformed this perception, revealing lipids as dynamic regulators of cellular function and disease.
The emergence of lipidomics, a major branch of systems biology, has enabled the large-scale identification, quantification, and functional analysis of thousands of lipid molecular species within biological systems. Unlike earlier decades when only a few hundred lipid molecules had been chemically characterized and cataloged, modern high-resolution analytical technologies—particularly mass spectrometry (MS) and liquid chromatography–mass spectrometry (LC-MS)—have uncovered enormous lipid diversity across cells, tissues, and organisms.
This expansion has elevated the lipidome, defined as the complete repertoire of lipids within a biological system, to a central component of modern omics research alongside genomics, transcriptomics, proteomics, and metabolomics.

The Lipidome as a Systems Biology Network
The lipidome is not simply a collection of hydrophobic molecules but an interconnected biochemical network integrated with metabolism, signal transduction, transcriptional regulation, and cellular homeostasis.
Lipids contribute to:
- Membrane architecture and fluidity
- Intracellular and extracellular signaling
- Energy storage and mobilization
- Inflammatory responses
- Cell growth and differentiation
- Apoptosis and programmed cell death
- Epigenetic and transcriptional regulation
- Disease pathogenesis and therapeutic response
In systems biology, lipids are now recognized as active components of metabolic and signaling networks rather than passive structural elements. Their interactions with proteins, nucleic acids, and metabolites influence emergent phenotypes including homeostasis, stress adaptation, development, and disease progression.
This systems-level perspective has driven the integration of lipidomics into computational biology and network medicine.
Rise of Bioinformatics in Lipid Research
The rapid growth of lipidomics generated a critical need for specialized bioinformatics frameworks capable of organizing, annotating, analyzing, and modeling lipid data.
Unlike nucleic acids and proteins, which are built from defined alphabets enabling straightforward sequence-based ontologies, lipids exhibit remarkable structural heterogeneity. Differences in carbon chain length, saturation, stereochemistry, functional groups, ring systems, and combinatorial isomerism create enormous complexity.
This complexity made traditional naming conventions based on common or historical names inadequate for large-scale computational analysis.
Lipid bioinformatics emerged to address major challenges including:
- Standardized lipid classification
- Universal nomenclature systems
- Computational structure representation
- Lipid databases and ontologies
- Pathway reconstruction
- Quantitative lipid network modeling
- Integration of lipidomics with genomics and proteomics
These developments laid the foundation for systems biology of the lipidome.
LIPID MAPS and the Evolution of Computational Lipidomics
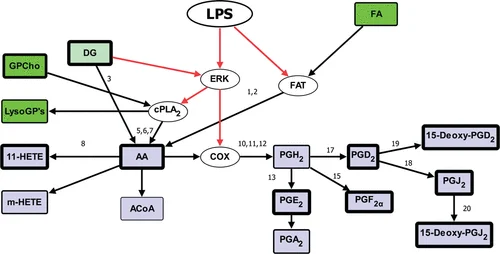
One of the most influential milestones in lipid bioinformatics has been the development of the LIPID MAPS initiative (Lipid Metabolites and Pathways Strategy).
LIPID MAPS was designed as a systems biology framework to characterize cell-wide lipid changes and reconstruct biochemical pathways involved in lipid metabolism and signaling.
Its goals include:
- Measuring global lipid responses to biological stimuli
- Mapping biosynthetic and metabolic pathways
- Integrating lipidomics with transcriptomics and proteomics
- Developing computational tools for lipid analysis
- Establishing standardized lipid classification systems
- Building searchable lipid databases
This initiative transformed lipidomics from descriptive chemistry into a computationally driven systems science.
Lipid Classification and Ontology
A foundational requirement for lipid bioinformatics is a robust classification and ontology system that is scalable, extensible, and computationally tractable.
The LIPID MAPS consortium introduced a chemistry-based classification system defining lipids as hydrophobic or amphipathic small molecules derived primarily from:
- Carbanion-based condensations of thioesters
- Carbocation-based condensations of isoprene units
Based on biosynthetic origin and structural principles, lipids are divided into eight major categories:
This category includes:
- Fatty acids
- Eicosanoids
- Fatty alcohols
- Oxylipins
- Fatty aldehydes
Fatty acyls function in energy metabolism, inflammation, signaling, and membrane composition.
Glycerolipids contain glycerol backbones esterified with fatty acids and include:
- Monoacylglycerols
- Diacylglycerols
- Triacylglycerols
These molecules are major energy storage lipids and signaling intermediates.
These are essential membrane components and include:
- Phosphatidylcholine
- Phosphatidylethanolamine
- Phosphatidylserine
- Cardiolipins
They participate in membrane structure and intracellular signaling.
Sphingolipids include:
- Ceramides
- Sphingomyelins
- Glycosphingolipids
They regulate apoptosis, stress responses, and membrane microdomains.
Prenol lipids include:
- Carotenoids
- Quinones
- Dolichols
These function in electron transport and cellular biosynthesis.
Polyketides include structurally diverse secondary metabolites with important pharmacological properties.
This hierarchical framework provides the foundation for lipid ontology and computational annotation.
Lipid Nomenclature and Standardization
Standardized nomenclature is critical for reproducibility and database interoperability.
The LIPID MAPS nomenclature system aligns with IUPAC-IUBMB principles while accommodating lipid-specific structural complexity.
It enables systematic naming based on:
- Carbon chain length
- Double bond position and geometry
- Functional group substitutions
- Positional stereochemistry
- Isomeric forms
For example: TG(16:0/18:1/18:2) : immediately communicates acyl composition for a triacylglycerol.
Such shorthand supports computational parsing and high-throughput lipid analysis.
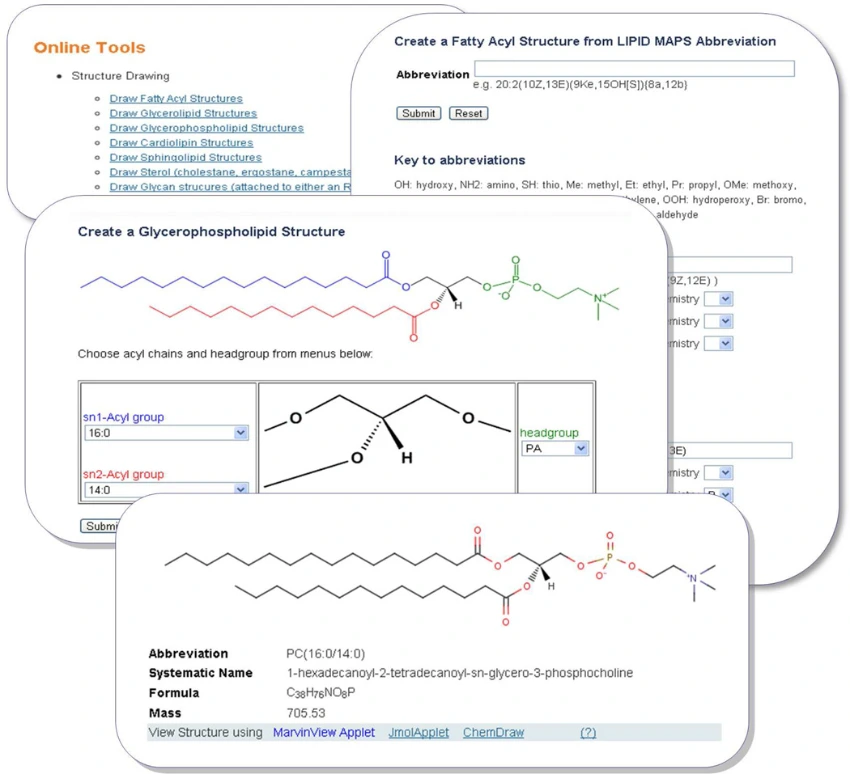
Automated Lipid Structure Drawing Tools
Manual drawing of complex lipids is labor-intensive and error-prone.
To address this, automated structure generation tools have been developed that:
- Generate structures from lipid abbreviations
- Produce MDL MOL and SDF files
- Assign systematic names automatically
- Calculate ontological descriptors
- Generate InChI and InChIKeys
These tools enable large-scale structure generation for tens of thousands of lipid molecules.
Important ontological descriptors generated include:
- Number of carbons
- Double bonds
- Rings
- Hydroxyl groups
- Amino groups
- Keto groups
- Molecular formula and mass
This automation has been transformative for database population and computational lipidomics.
Lipid Databases in Bioinformatics
Lipidomics depends heavily on specialized databases for storage, retrieval, annotation, and systems integration.
LIPID MAPS Structure Database (LMSD)
LMSD is one of the most comprehensive dedicated lipid databases.
It contains:
- Over 30,000 curated lipid structures
- Classification hierarchies
- Ontological descriptors
- Cross-references to external resources
- InChI identifiers
- Exact masses and formulas
- Structural search tools
Conclusion
Bioinformatics and systems biology have transformed lipid research from descriptive chemistry into an integrative computational science. Advances in lipid classification, ontology, structural representation, databases, pathway modeling, and large-scale data analysis have established the lipidome as a core component of modern systems biology.
Through initiatives such as LIPID MAPS and the development of computational lipidomics tools, researchers can now analyze lipid diversity, map lipid networks, model metabolic fluxes, and integrate lipid data into broader biological frameworks.
As lipidomics continues to merge with genomics, proteomics, metabolomics, and artificial intelligence, it is poised to become one of the most powerful approaches for understanding cellular regulation, disease mechanisms, and future precision medicine.
Commencez à écrire ici ...